Cancer organoids: an important new tool in the armoury of cancer biologists
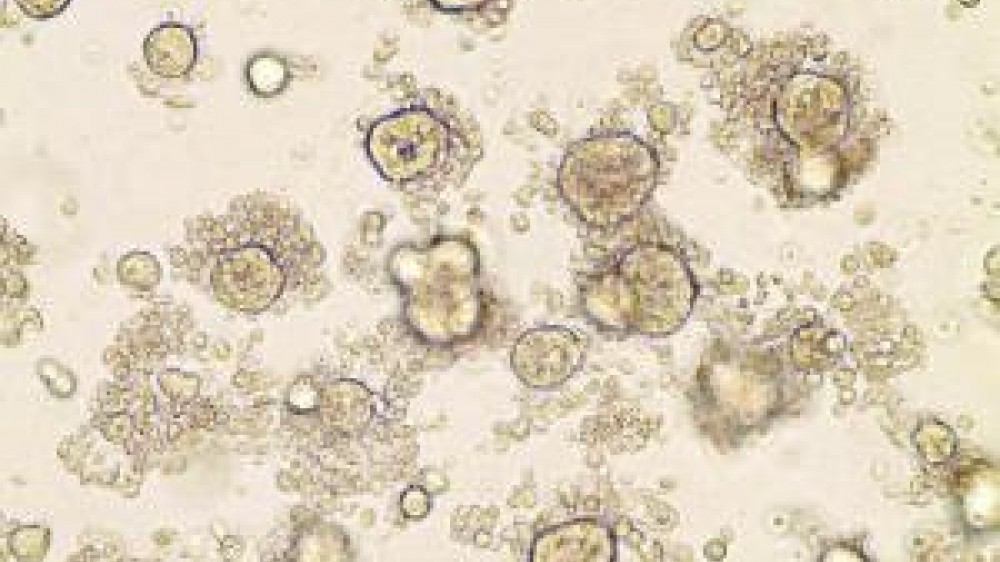
A study, published in the journal Cell, describes organoids of colorectal cancer derived from the patients’ own tissue.
The creation of organoids, 3D organ-buds grown in vitro that show realistic micro-anatomy, is a technique being investigated across a huge number of different cell types, from the brain to the gut.
The colorectal cancer collection (‘biobank’) described in the study was developed by Dr Hayley Francies from the Wellcome Trust Sanger Institute and her colleagues. Earlier this year, Dr Francies received a highly commended award for this work in our annual 3Rs Prize competition. The award panel was impressed with the work and its potential use in high throughput drug screening in cancer research, therefore reducing the reliance on animal models.
Cancers, including those of the colon and rectum, have many subtypes which display different combinations of genetic alterations. This diversity is important to understand in patients: cancer drugs specifically target different cellular pathways, so depending on the genetic background of the person and the tumour, patients’ response to therapy can vary. Cancer treatments are therefore becoming more targeted to specific patients, and effectively ‘personalised’ for each person. Dr Francies and her colleagues propose their collection of organoids as an improved, non-animal model to help meet the goal of a more personalised approach.
Bridging the gap between cell lines and animals
Systems currently used to study cancer range from simple 2D cell cultures, which lack the necessary structural and functional complexity of the original tumour, to patient-derived xenograft (PDX) mouse models. To create PDX models, fragments of patients’ tumours are implanted under the skin of immune-deficient mice. The efficacy of potential anticancer reagents is often assessed in PDX mice, and current methods mean that testing one compound can require in the region of 50 animals.
Dr Francies said: “Mouse models are the standard at the moment, but they are time-consuming and costly, and the animals can experience pain and distress because of tumour growth. Even though we are unable to completely replace mouse models at this stage, we aimed to make a bank of better non-animal cancer models, to limit the use of animals in total. We sought to create a model that would allow us to study colorectal cancer easily in vitro while capturing a level of complexity not found in cell lines.
“Like PDX mice, organoids are personalised because they are created from patient’s own tissue. Organoid culture can be easily propagated for high throughput studies. This way, many potential anticancer compounds can be screened and a more informed decision can be made – to test the same number of drugs in an animal model would never be feasible.”
The fact that these organoids can be easily manipulated, and various endpoints can be measured, means that they also have applications for basic research, where researchers can observe intricate molecular changes that would be more difficult to observe in mice.

Dr Francies talks about her work to a guest at the 3Rs Prize ceremony. (NC3Rs)
How does it work?
To create the organoids, the team at the Sanger Institute worked together with colleagues at the Broad Institute of MIT and Harvard, USA, and the group of Professor Hans Clevers from the Hubrecht institute, The Netherlands. The researchers took sections from patients’ colorectal tumours and healthy surrounding tissue from the colon epithelium. They analysed the DNA sequence of the sections, to look for cancer-related mutations in the patient’s cells. Simultaneously, the researchers used the patient’s tissues to start cultures of both tumour and normal organoids. With the use of newly adapted culture conditions for long-term expansion in 3D, they observed ~90% success rate, managing to grow 19 normal and 22 tumour organoid cultures from 20 patients. Normal organoids showed organisation similar to the original tissue, with different cell types found in the healthy colon, while the structure in cancer organoids was completely lost, and the cultures between different patients showed a lot of variety.
In the second stage of the study, Dr Francies and her colleagues analysed the drug response of the organoids by screening them against a library of 83 compounds, including drugs in clinical use, chemotherapeutics, and drugs previously investigated or undergoing studies in clinical trials, as well as other compounds aimed at a diverse range of cancer targets. The tumour organoids were tested against this library in a robotised screen, generating around 5,000 measurements of organoid-drug interactions. In comparison, this information would have been obtained with the use of at least 5,000 mice, signalling a large reduction in the use of animals. The screen identified the gene-drug associations already known and reported in literature, as well as some possible new interactions.
Chief Executive of the NC3Rs, Dr Vicky Robinson said: “Collectively, these results indicate that organoids have the ability to accurately reflect the molecular features and heterogeneity of cancer. Growing 3D organoid cultures from patient tumours could transform cancer research by better reflecting the biology and therapeutic response of patient tumours, while reducing the use of mouse models. The use of organoids provides an unprecedented opportunity to evaluate the effectiveness of cancer drugs in a more accurate way and at scale not possible in PDX models.”
What’s next?
When asked about the 3Rs Prize, Dr Francies commented: “Given the potential of cancer organoids to identify novel cancer therapies and biomarkers of drug response, we at the Sanger Institute want to develop large collection of organoid models which will allow us to reduce the use of animals in cancer research. The Prize funds will help explore another aspect of cancer research: drug resistance, a phenomenon often seen in the clinic but not addressed by many drug screens.
“Using the funds from the Prize, we intend to scale up the existing drug screen for testing combination drug therapies and assess numerous different, clinically-relevant drug combinations. To do that, the screening throughput will have to increase in capacity which would not be plausible in animals (one 1536-well plates screening can assess ~ 200 drug combinations).”
The international NC3Rs 3Rs Prize, sponsored by GlaxoSmithKline, is awarded every year to highlight recent publications with exceptional contribution to scientific and technological advances in the 3Rs in medical, biological or veterinary sciences, published in a peer-reviewed journal in the last three years. The prize consists of a grant of £18k, plus a personal award of £2k and highly-commended entries receive a £4k grant and £1k personal award. The applications are judged by an independent panel of experts and the prize is presented during an award ceremony, attended this year by more than 150 NC3Rs stakeholders and guests from the scientific community.
Two joint main prizes last year were given to researchers looking at ways of improving welfare of laboratory dogs, and a method to develop for the first time organoids reflecting the embryonic human brain. Nominations for 3Rs Prize 2016 will open in the autumn, so watch this space!
