Supporting the adoption of microphysiological systems for COVID research
At a glance
Contents

Overview
We have established a working group with the US National Toxicology Program Interagency Center for the Evaluation of Alternative Toxicological Methods (NICEATM). This group will help coordinate global efforts to use microphysiological systems (organ-on-a-chip and other complex multicellular in vitro models) reducing the reliance on in vivo studies of COVID-19 and future infectious diseases.
The COVID-19 pandemic has resulted in a significant number of animal models including mice, hamsters, ferrets and non-human primates, to better understand disease biology, and develop and test treatments and vaccines. However, the variability in susceptibility and manifestation of symptoms, which can differ substantially from humans, has emphasised the need for models based on human biology.
Human-based microphysiological systems (MPS) of the lungs and other key organ systems affected by COVID-19 are therefore rapidly being developed. Such a rapid response on a global scale risks fragmentation and resource duplication. This presents significant challenges for model developers, drug/vaccine manufacturers and regulatory authorities in coordinating efforts and understanding the utility and validity of the new systems.
To support the biosciences research community in overcoming these challenges, the working group will address the following objectives:
- Provide a neutral forum to facilitate interaction and engagement between international collaborative research efforts.
- Raise awareness of COVID-19 MPS technologies and support their application in assessing the safety and efficacy of potential novel therapeutics through building connections between technology developers and end-users.
- Work with global authorities to understand how MPS models can be considered in a regulatory context.
- Provide cross-discipline and -sector expertise in discussing and characterising model performance and readiness criteria.
- Support the assessment of MPS against gold-standard preclinical (in vivo) and clinical data.
- Ensure the 3Rs opportunities these model platforms offer are recognised.
This working group is in partnership with the US National Institute of Allergy and Infectious Diseases, Division of Microbiology and Infectious Diseases, US Army DEVCOM Chemical Biological Center, and US National Center for Advancing Translational Sciences (NCATS). Members include researchers, MPS model developers, therapeutic/vaccine manufacturers, and international regulators.
First working group meeting: Recording available
The MPSCoRe working group held its first meeting 29 January 2021. Among the topics discussed were the limitations of animal models for studying COVID-19 and the need for high-quality study data to evaluate MPS technologies for COVID-19 studies. A video and summary of the meeting presentations can be viewed on the U.S. National Toxicology Program website.
If you are interested in finding out more about this project, please email us with your name, role and area of expertise.
Programme of work
Supporting the development of a COVID-19 disease portal in the Microphysiology Systems Database
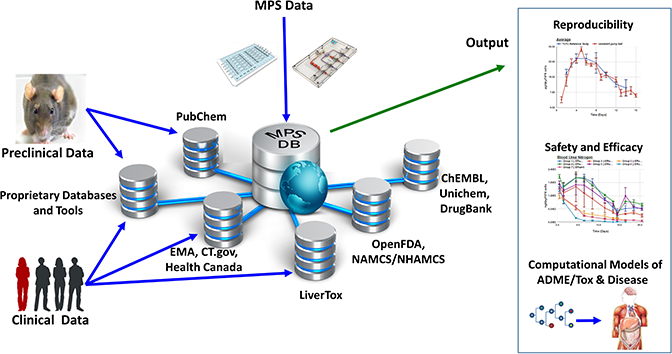
The Microphysiology Systems Database (MPS-Db) is a web-based resource supported by NCATS to accelerate the development and application of MPS for basic research and drug development. It collates and makes accessible experimental data submitted by MPS model developers and preclinical and clinical data from a range of open source and proprietary databases including PubChem, CheMBL, Genbank, PubMed, ClinicalTrials, OpenFDA and EMA. The database includes functionality to manage, analyse, share, computationally model and integrate data in one platform, as well as tools to assess the reproducibility and transferability of MPS experimental models.
NICEATM is partnering with the MPS-Db developers to expand the platform to include a COVID-19 disease portal to accelerate the development and adoption of human MPS for testing prospective therapeutics, a key aim of the MPSCoRe working group. The portal will exist within the framework of the existing MPS-Db and will include:
- Summaries and links to experimental model designs and study components including reagents, cells, devices, compounds, assays and assay kits, which can be used to develop COVID-19 in vitro MPS models.
- Links to in vivo and computational model data of COVID-19 and databases of clinical trial data that could inform the design, implementation, and validation of MPS models (e.g. the National Covid Cohort Collaborative Data Enclave).
- User guides to assist with the addition of models, creation of studies, uploading data, and analysis of results.
- A permissions structure that will allow users to choose when their data can be viewed by the MPSCoRe group and when it can be released publicly.
Progress on the development of the COVID-19 disease portal will be reviewed by the MPSCoRe working group and will be reported on this webpage. Additional programmes of work will also be added as they are agreed and initiated.
The MPSCoRe hosted a webinar to provide an in depth tutorial on how to use the MPS-Db to find and share data on COVID-19 projects. The webinar recording can be accessed on the U.S. National Toxicology Program website.
Stakeholder engagement workshop
A workshop of the MPSCoRe was held in April 2021 to engage relevant stakeholder groups in the project, to share project outputs and priorities and to showcase the latest advances in the development and application of MPS for infectious disease research and therapeutic development. The workshop also provided an opportunity for the community to feedback their views on the working group programme of work and help shape the project outcomes.
A recording of the workshop can be viewed on the U.S. National Toxicology Program website.
Supporting access to COVID-19 variants of concern
Working with the World Health Organisation we have integrated MPSCoRe member research capabilities into global efforts to study Omicron and other variants of concern and to advance the development of COVID-19 medical countermeasures (vaccines, therapeutics and/or drugs). This includes supporting MPSCoRe members in accessing Omicron viral samples and other materials from global suppliers such as BEI Resources and the WHO BioHub so that MPS and animal studies can be delivered concurrently and enable timely comparison of the utility and validity of different model systems to address specific research questions.
Publications
- Kleinstreuer N, Holmes A. 2021. Harnessing the power of microphysiological systems for COVID-19 research. Drug Discov Today 26(11):2496-2501. doi: 10.1016/j.drudis.2021.06.020.
