Wax moths and tuberculosis infection
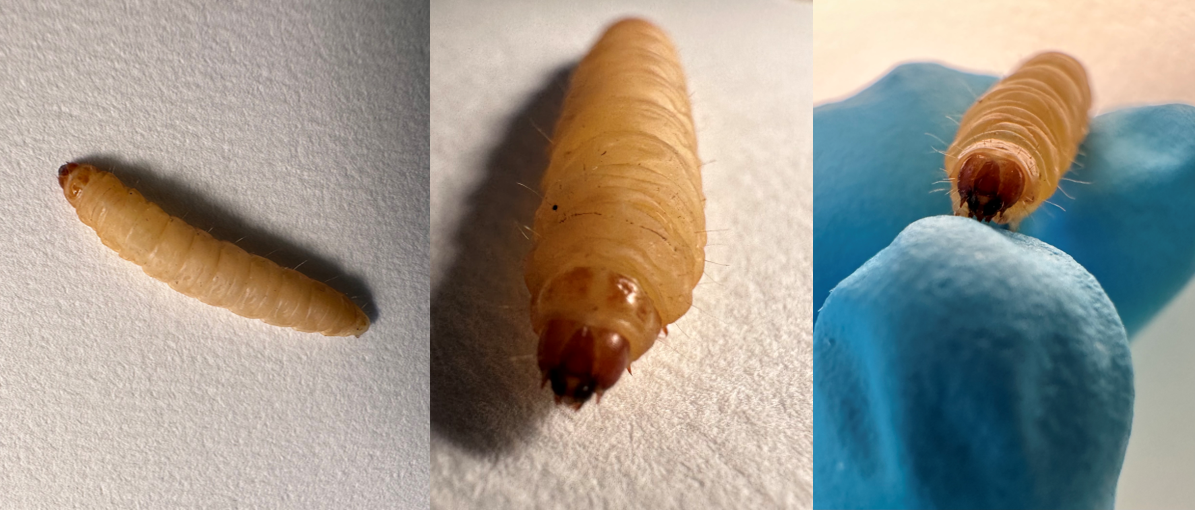
Professor Paul Langford and colleagues Drs Brian Robertson, Sandra Newton and Yanwen Li were awarded funding to develop a tuberculosis infection model using Galleria mellonella to replace the use of mammals in some studies.
Tuberculosis (TB) is a major cause of death worldwide with over 10 million people infected each year. Infections are caused by the bacterium Mycobacterium tuberculosis and can be treated with antibiotics. However, there is increasing incidence of multi-drug resistant TB infections and there is a focus in TB research on identifying new compounds for targeting drug-resistant TB. A range of animal models are used including mice, guinea pigs, rabbits and monkeys but no animal model replicates all aspects of TB pathogenesis. Mice are the most commonly used and show some disease symptoms such as raised temperature, respiratory distress and weight loss. Depending on the number of doses being tested each drug screening experiment can require up to 200 mice. Mice do not however develop granulomas, aggregates of immune cells formed to prevent bacteria from spreading further, a key characteristic of human TB infections.
3Rs benefits
In 2018, Paul and colleagues published a proof-of-concept study demonstrating Galleria mellonella (the wax moth) could be infected with mycobacteria. With further funding from the NC3Rs, Paul and student Masanori Asai established and characterised a Galleria model of TB infection, which possessed hallmark features of human TB disease, including granuloma-like structures. Masanori has six publications from his PhD studies that demonstrate the utility of the Galleria model in both infection studies and drug screens. A methods article in the Journal of Visual Experiments describing key practical details about how to use Galleria in TB research, including a video of the method, has been viewed nearly 8,000 times.
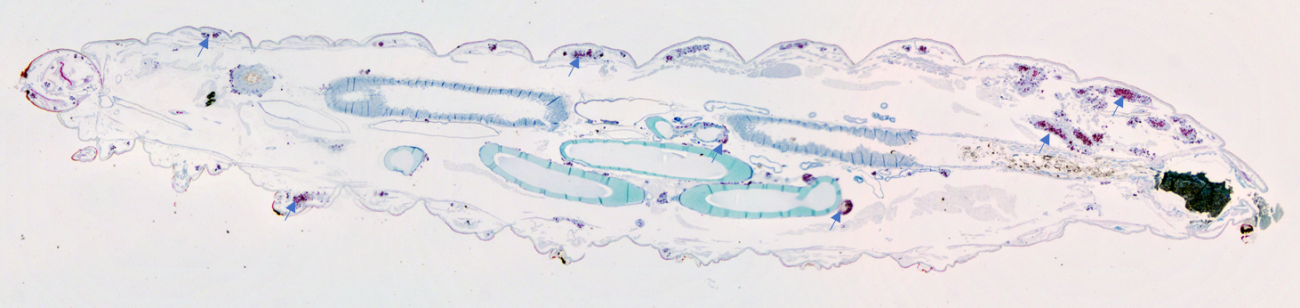
Scientific and technological benefits
Mycobacterial strains with varying virulence are commonly used in TB research with the most virulent being used in containment level 3 laboratories because of the biosafety concerns. Masanori has challenged Galleria with mycobacteria at high containment and shown that the model can discriminate virulence between different strains and mutants. Identifying mycobacterial genes that are present in virulent strains and when knocked out lead to loss of virulence is important as the products of these genes are potential targets for novel antibiotic targets.
A further advantage is that Galleria larvae can be maintained and infected at 37oC allowing studies to be performed at the same temperature as TB infections in humans. This is particularly important for drug screens as temperature can affect mycobacterial physiology. Screening drug efficacy in Galleria takes approximately four to five days while testing in mice can take many weeks. Masanori screened five antibiotics currently used to treat TB, including both first-line therapies and second-line that are used for infections resistant to first-line treatments. Isoniazid and rifampicin showed the most efficacy in Galleria infections, which is comparable to studies in mice. Publications and presentations of the work at conferences has led to new collaborations testing novel antibiotics in Galleria for the treatment of multi-drug resistant TB.

Added value
Masanori has subsequently been awarded an NC3Rs Training Fellowship to further characterise TB infections in Galleria, specifically the granuloma-like structures confirming their similarity to human granulomas building confidence in the model as an alternative to rodent models. A major barrier for Galleria research is the lack of reagents to detect specific immunological markers, such as inflammatory markers, and Masanori is addressing this as part of his Fellowship by analysing transcriptome data, and developing RNA probes and inhibitors. Imperial College London has awarded Masanori the Provost’s Award for Excellence in Animal Research recognising his work replacing mammalian animals in TB research.
Further details of this PhD Studentship including application abstract and publications.

